Over the last decade, the field of metagenomics has grown extensively, leading to an exponential increase in sequencing data in public data repositories. Availability of this huge collection of data has accelerated the discovery of novel microbiome-related biomarkers and microbiome-based novel biomedical therapeutics. One of the key steps in identifying potential biomarkers or lead candidates is the availability of relevant clinical and experimental metadata associated with each sample or specimen. Annotation of metadata includes fully describing the sample type, procedure of collection, extraction and assay methods, and relevant clinical phenotypes. Lack of complete annotations/metadata may negatively impact downstream statistical steps and follow-up studies aiming to reuse the data.
Our goal at CosmosID is to make it easy and seamless for our users to leverage their metadata for gaining actionable insights from their microbiome data. Therefore, we are pleased to announce a new metadata dashboard as an additional feature on the CosmosID-HUB Microbiome platform. This feature will allow users to not only upload their metadata for their associated samples easily, but also to search and make comparative analyses automatically with predefined cohorts using user-uploaded metadata. The CosmosID-HUB Microbiome will provide an infrastructure where a user can go from raw data to translational or actionable insights within a few hours of uploading their data. The ability to associate metadata with raw sequencing data will also allow users to leverage CosmosID-HUB as a centralized and secure storage hub for all their projects.
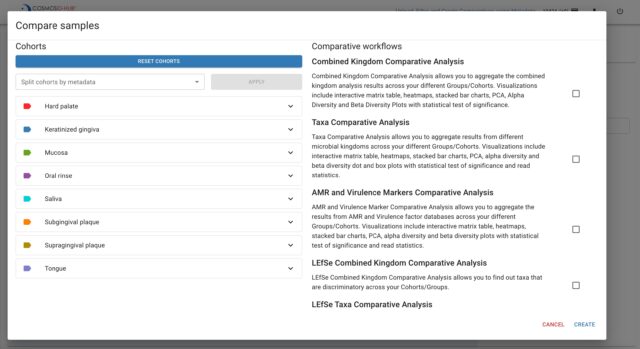
The new release of the Metadata feature is part of CosmosID’s microbial genomics platform, CosmosID-HUB Microbiome, which is continually expanding with various features and workflows for analyzing microbial communities and for inferring translational and actionable microbiome insights. Currently, CosmosID-HUB Microbiome analyzes 16S, whole genome (shotgun metagenomics and metatranscriptomics), and ITS sequencing data and provides options for discovering antimicrobial resistance (AMR) and virulence markers, microbial taxa (composition), and functions. Users are also able to aggregate and compare their primary analysis results (taxa, AMR and virulence markers, functions) by cohorts or groups using statistical modules to gain actionable insights from their data. The overall CosmosID-HUB platform also includes our COVID-19 pipeline, which detects the presence of COVID-19 and characterizes variants of concern or interest in both clinical and wastewater specimens. Researchers are provided the flexibility to access and pay for the type of analysis they require based on their research question.
Interested in trying out the metadata dashboard? Set up your free HUB account here!
If you are interested in more details or a live demo of the CosmosID-HUB, reach out to us at sales@cosmosid.com!


